Let our intelligence move you
However you want to change the world, we’ll get you there.

Dig deeper into your field. Collaborate with the right partners. Build a reputation for excellence.
We’ll give you a single source of information to help your institution lead.

Get right to real-life outcomes, with patients at the heart of everything you do.
We’ll help you collect and analyze information for insights you can easily act on.
Go head-to-head with the competition, and win.
We’ll help you stand out, build a reputation for unbeatable client service, and streamline your processes.

Go to market faster. Protect and manage Intellectual Property. Free up your team to focus on strategic outcomes.
We’ll give you the tools and services you need to make informed decisions at speed.
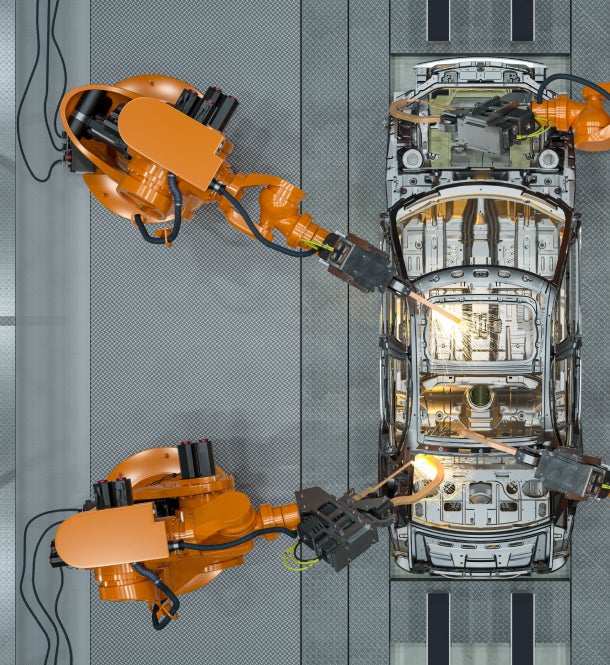

How Gedeon Richter partnered with Clarivate consultants to transform regulatory workflow